Project Summary
CRISPR Diversity and Antimicrobial Susceptibility of Salmonella Isolates from Dairy Farm Environments in Texas
- Principle Investigator(s):
- Kevin Cummings1, Tom Edrington2, Guy Loneragan3
- Institution(s):
- 1Texas A&M University
- 2USDA Agricultural Research Service (ARS)
- 3Texas Tech University
- Completion Date:
- June 2014
Background
Dairy cattle are a key reservoir of several Salmonella serovars that are leading causes of human salmonellosis, including multidrug‐resistant (MDR) S. enterica serovar Newport and S. enterica serovar Typhimurium. Administration of antimicrobial agents to dairy cattle and other food animals is considered to be one of the driving factors for antimicrobial resistance among Salmonella and other pathogens, based on exertion of local selection pressure in agricultural settings. Antimicrobial agents that are currently licensed for use in dairy cattle in the United States include enrofloxacin, florfenicol, and various penicillins, cephalosporins, macrolides, sulfonamides, and tetracyclines; extra‐label use of some additional drugs is also permitted under certain circumstances. However, a number of studies have failed to find conclusive evidence that the use of antimicrobial agents in animal production systems leads to a sustained increase in antimicrobial resistance among pathogens within the gastrointestinal tract of animals. In fact, recent experimental work suggests that the environment could play a central role in antimicrobial selection pressure. Excreted ceftiofur metabolites were found to exert substantial selection pressure that promoted resistance in the environment, and cattle subsequently became infected with resistant bacterial strains through environmental exposure. This suggests that there is likely to be a discrepancy among environmental reservoirs in their ability to promote the emergence and persistence of antimicrobial resistance on dairy farms. Identification of high‐risk environmental reservoirs within dairies would provide an opportunity to mitigate the burden of antimicrobial resistance at the pre‐harvest level.
The objectives of the study were to: 1) determine the prevalence and diversity of Salmonella in multiple environmental locations within dairy farms throughout Texas, 2) characterize the antimicrobial susceptibility status of Salmonella isolates across dairy farm environmental locations, and 3) establish the degree to which environmental contamination reflects carriage of resistant Salmonella among cull dairy cattle.
Methodology
Data for this study were collected prospectively from a convenience sample of dairy farms throughout Texas. Following enrollment, dairy farms were sampled during the period from August – October 2013 using a cross‐sectional approach. Eight samples were collected from each of four environmental locations per farm for Salmonella culture: hospital pen, maternity pen, cow housing area, and calf housing area. Fecal samples were also collected from cows due to be culled from each of the enrolled dairies, as available at the time of environmental sampling; these were likewise cultured for Salmonella. Standard bacteriologic culture methods were used to isolate Salmonella from samples. Clustered regularly interspaced short palindromic repeats (CRISPR) subtyping and antimicrobial susceptibility testing was performed on all isolates. Data were analyzed to characterize the diversity and antimicrobial resistance of Salmonella isolates across dairy farms and specific environmental reservoirs.
Findings
Salmonella was isolated from 67.0% (236/352) of environmental samples and 64.2% (43/67) of cull cow fecal samples. Prevalence of Salmonella in the environment varied widely among the 11 farms, ranging from 31.3% to 96.9%. Salmonella prevalence varied significantly by environmental location (Fig. 1), with higher proportions of positive samples from the maternity (83.0%, 73/88) and hospital pens (68.2%, 60/88) and lower proportions from the calf housing (58.0%, 51/88) and cow housing areas (59.1%, 52/88).
Preliminary results indicate a tremendous diversity of Salmonella subtypes within dairy farm environments. Degree of diversity appears to vary across environmental locations.
Resistance to individual antimicrobial agents was generally low, ranging from 0% to 11.6% of all isolates tested (Table 1). However, some isolates were resistant to as many as 11 drugs. Multidrug resistance, defined here as resistance to two or more classes of antimicrobial agent, was evident in 11.9% (27/226) of environmental isolates and 19.5% (8/41) of fecal isolates. Five (45.5%) of the farms in this study yielded MDR Salmonella isolates from the environment, cull cattle, or both. Isolates from the calf housing area and maternity pen were significantly more likely to be MDR than isolates from the cow housing area and hospital pen.
Implications
Salmonella was ubiquitous in dairy farm environments, with a discrepancy in prevalence across environmental locations. The calf housing area and maternity pen were more likely to yield multidrug‐resistant Salmonella isolates than other environmental reservoirs. Targeted environmental hygiene measures might provide a cost‐effective opportunity to mitigate the burden of antimicrobial resistance at the pre‐harvest level. The high diversity of Salmonella subtypes within dairy farm environments will facilitate future work to investigate subtype persistence over time and/or subtype turnover.
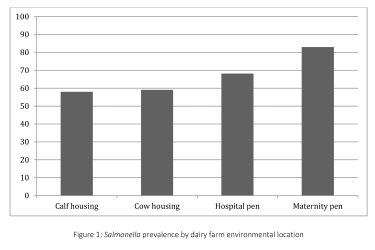
Table 1: Resistance to individual antimicrobial agents among 267 Salmonella isolates obtained from dairy farm environments and cull cattle in Texas, 2013.
|
Antimicrobial agent |
Resistance, % |
|
Tetracycline |
11.6 |
|
Sulfisoxazole |
10.9 |
|
Amoxicillin/clavulanic acid |
7.9 |
|
Ampicillin |
7.9 |
|
Ceftiofur |
7.9 |
|
Ceftriaxone |
7.9 |
|
Streptomycin |
7.5 |
|
Chloramphenicol |
6.4 |
|
Cefoxitin |
5.6 |
|
Trimethoprim/sulfamethoxazole |
2.2 |
|
Kanamycin |
1.1 |
|
Azithromycin |
0 |
|
Ciprofloxacin |
0 |
|
Gentamicin |
0 |
|
Nalidixic acid |
0 |